500% more leads. 35% less marketing spend.
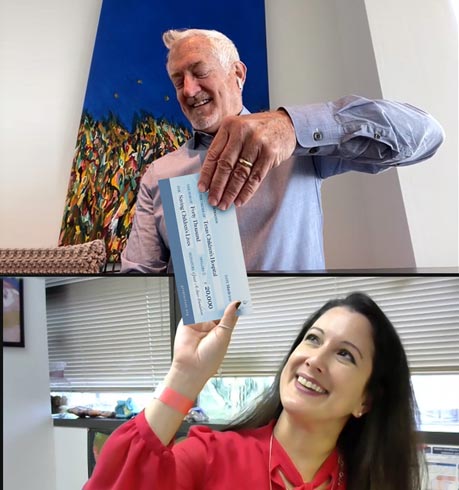
HexaGroup has been supporting this wonderful project for over 10 years and we are proud of the foundation’s achievements in advancing the field of pediatric diagnostic testing.
The Grant-A-Starr Foundation has contributed nearly $1.6 million to world-class projects to advance diagnostic testing to save children’s lives.
Our dedicated donors can be proud to be part of a revolution to transform diagnostic testing.
Same Mission, New Strategic Direction
Tremendous strides have been made in the last decade to advance the diagnostic testing methods that identify infectious diseases. Case in point – the term ‘metagenomics’ was only recently defined in 2005 by two researchers at the University of California, Berkeley.
This new molecular way of analyzing microbial pathogens that make us sick (e.g. bacteria, viruses) has opened up fresh perspectives to diagnose illnesses more quickly and more accurately. The Grant-A-Starr Foundation is honored to be a part of this scientific revolution.
As the medical community embarks on a new era of diagnostic testing capabilities, the Grant-A-Starr Foundation sets forth in a new strategic direction with a move away from fundraising to information sharing. While the foundation will no longer solicit donations, we will continue to share information about advancements resulting from the great work underway at the exceptional organizations we supported and collaborated with over the years – Texas Children’s Hospital, Children’s Memorial Hermann Hospital and IDbyDNA.
From the bottom of our hearts – thank you to all the donors, volunteers and supporters who continue to help advance diagnostic testing and celebrate Grant’s life.
About Grant-A-Starr Foundation

Grant's Story
Grant’s life was shortened by an unknown childhood infection. We celebrate Grant’s life with the Grant-A-Starr Foundation with the mission to advance diagnostic testing for infectious disease to help others.
Early Diagnosis
Grant-A-Starr Foundation funds the research and development to diagnose illnesses more quickly and accurately.
Our Research Partners
The Grant-A-Starr Foundation contributed nearly $1,600,000 towards the early diagnosis of infectious disease.
Texas Children’s Hospital
Advancing molecular diagnostics development in a pediatric setting to shorten the time required to detect dangerous bacterial and viral pathogens.
Children’s Memorial Hermann Hospital
Establish a simple and cost-effective algorithm for early diagnosis and effective management of Aseptic Meningoencephalitis in children and adults.




